 
The settings used to call the consensus are set under the Display tab to the right of the sequence viewer. This is the consensus of the reads only and does not include the reference sequence. Zoom in so that you can see the sequence bases. At the top of the contig you’ll see a consensus sequence. You can see that most pairs map with inserts of 450-500 bp, close to the expected size of 500 bp. Click on the Insert sizes tab above the sequence viewer to see the actual distribution of insert sizes. In this contig you can see that most of the reads are green, meaning that they map with approximately the expected insert size. This setting allows you to see at a glance whether your paired-end reads map at the expected distance apart, based on the insert size you specified when you set up the paired reads. Now go back to the General tab and ensure that Colors is set on “Paired Distance”. This will display the reads in horizontal rows. 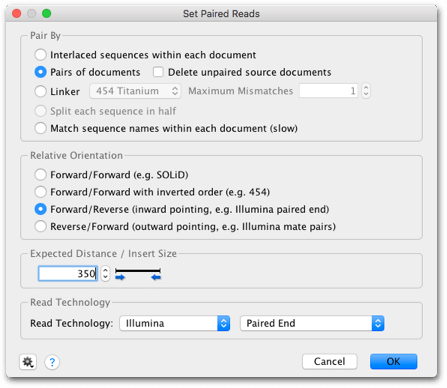
Under the Advanced settings tab to the right of the sequence viewer, ensure that “Vertically compress contig” is checked. Open the contig document (which should be called “yghJ paired Illumina reads (trimmed) assembled to yghJ CDS (divergent reference)”) to see how the reads map to the reference sequence. Your Map to Reference options should now look like this: Uncheck “Save in subfolder” if it is selected. The Fine Tuning option can improve the results by aligning reads to each other in addition to the reference sequence – set this on “Iterate up to 5 times”.Įnsure Do not trim is selected, and under the Results panel, choose Save assembly report and Save contigs. For next-generation sequencing, Medium or Medium-Low sensitivity is recommended, as using High Sensitivity will take a long time and is unlikely to improve the results if you have sufficient coverage. Geneious Prime will automatically choose an appropriate sensitivity based on the size of your dataset. The Sensitivity should be set on Medium Sensitivity/Fast. A brief outline of the different algorithms available is given here. A number of different mapping algorithms are available here, and these have varying strengths and weaknesses depending on the type of data you are assembling. In the Method panel, ensure Geneious is chosen as the Mapper. Instead, it can be selected from any folder in your database by clicking the Choose button within the setup options. 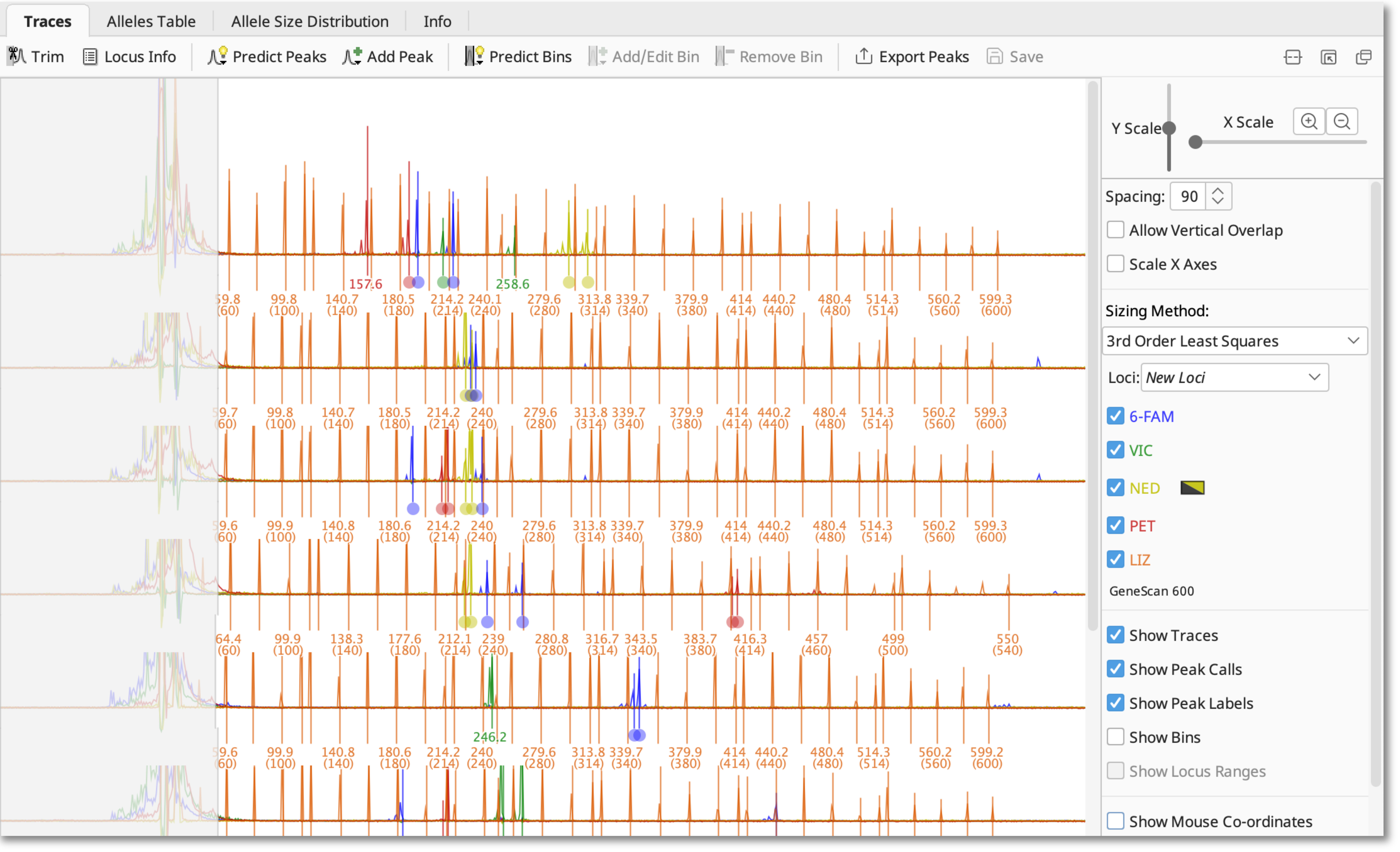
Note that in Geneious 8.1 and above, the reference sequence does not have to be pre-selected before opening the Map to Reference window. In this case it has chosen correctly, and “yghJ CDS (divergent reference)” should be shown as the reference sequence. Geneious Prime will guess which of the sequences you have selected is the reference, but you can change this using the drop-down menu. Click Align/Assemble → Map to Reference, and reset to the default setting using the settings cog at the bottom left of the window. Holding down the shift key, select your file of trimmed reads, and the reference sequence (yghJ CDS ). 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
